CHANNELS

Michael Ignarski, PhD
+49 221 478-84176
michael.ignarski@uk-koeln.de
TEAM MEMBERS
Steffen Hinrichs
Bernhard Wagener
Michael Ignarski
RNA-biology and metabolism in kidney disease
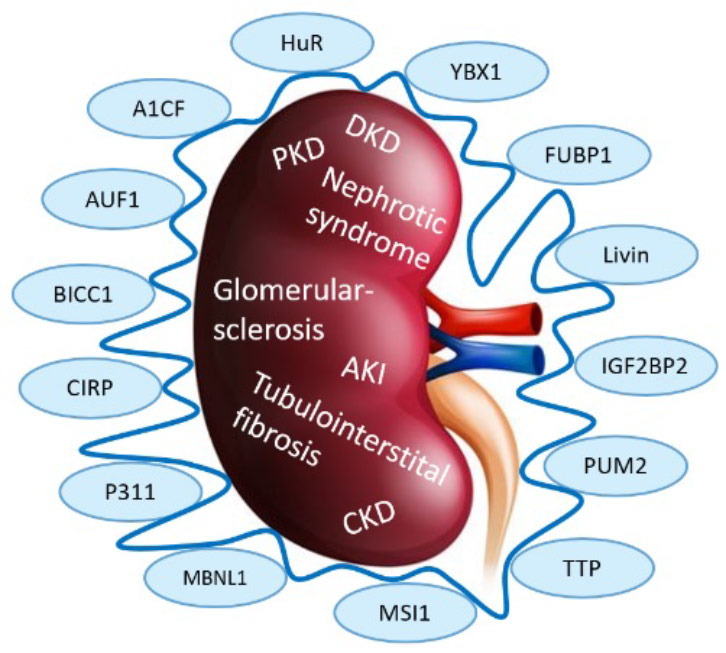
RNA binding proteins (RBPs) play critical roles in RNA biology and metabolism, and their dysregulation contributes to the pathogenesis and progression of kidney disease. The kidneys are essential organs responsible for maintaining fluid and electrolyte balance, regulating blood pressure, and eliminating waste products. Disruptions in RNA-related processes, including transcription, splicing, and translation, can have profound effects on kidney function.
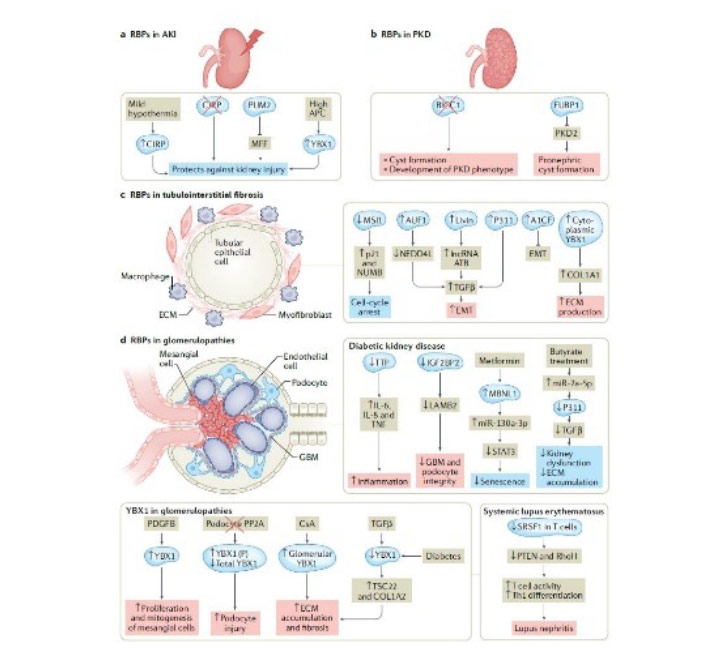
RBPs are key regulators of post-transcriptional RNA processing and function by interacting with RNA molecules to control their stability, localization, and translation. Dysregulated expression or activity of RBPs can lead to altered RNA processing, resulting in aberrant gene expression and cellular dysfunction. In kidney disease, dysregulation of RBPs has been implicated in various pathological conditions, such as glomerular diseases, polycystic kidney disease and acute kidney injury.
Furthermore, RBPs have emerged as critical players in the regulation of metabolic processes. They can modulate the expression and activity of enzymes involved in metabolic pathways, including glucose and lipid metabolism. Dysregulation of these metabolic pathways can contribute to cellular stress, inflammation, and fibrosis, ultimately leading to kidney dysfunction.
Recent advancements in high-throughput sequencing technologies and computational approaches have allowed comprehensive profiling of RBPs and their RNA targets in kidney disease models and patient samples. Integration of multi-omics data, including genomics, transcriptomics, and proteomics, has provided valuable insights into the complex interplay between RBPs, RNA biology, and metabolism in kidney disease.
Further research is needed to elucidate the specific mechanisms by which RBPs contribute to metabolic changes and kidney disease and to explore their potential as therapeutic targets for intervention in renal disorders.

SELECTED PUBLICATIONS
Seufert L, Benzing T, Ignarski M, Müller RU. RNA-binding proteins and their role in kidney disease. Nat Rev Nephrol. 2022 Mar;18(3):153-170. doi: 10.1038/s41581-021-00497-1.
Talyan S, Filipów S, Ignarski M, Smieszek M, Chen H, Kühne L, Butt L, Göbel H, Hoyer-Allo KJR, Koehler FC, Altmüller J, Brinkkötter P, Schermer B, Benzing T, Kann M, Müller RU, Dieterich C. CALINCA-A Novel Pipeline for the Identification of lncRNAs in Podocyte Disease. Cells. 2021 Mar 20;10(3):692. doi: 10.3390/cells10030692.
Johnsen M, Kubacki T, Yeroslaviz A, Späth MR, Mörsdorf J, Göbel H, Bohl K, Ignarski M, Meharg C, Habermann B, Altmüller J, Beyer A, Benzing T, Schermer B, Burst V, Müller RU. The Integrated RNA Landscape of Renal Preconditioning against Ischemia-Reperfusion Injury. J Am Soc Nephrol. 2020 Apr;31(4):716-730. doi: 10.1681/ASN.2019050534.
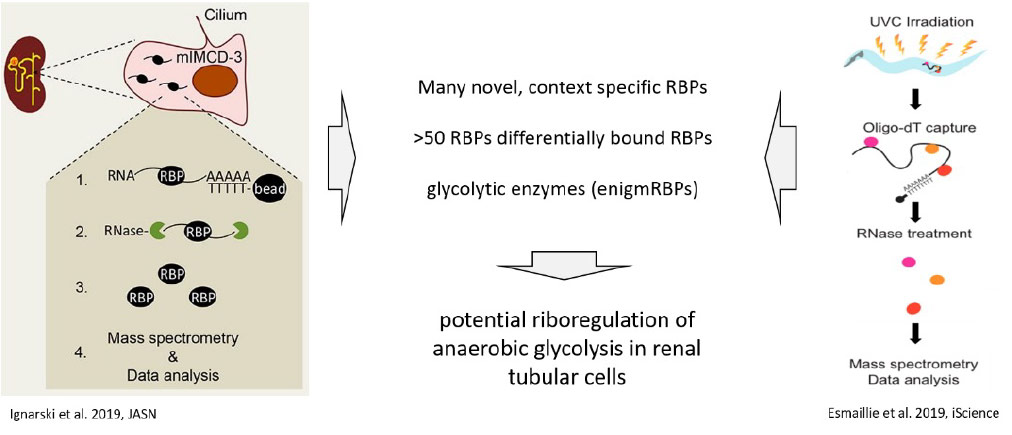
Esmaillie R, Ignarski M, Bohl K, Krüger T, Ahmad D, Seufert L, Schermer B, Benzing T, Müller RU, Fabretti F. Activation of Hypoxia-Inducible Factor Signaling Modulates the RNA Protein Interactome in Caenorhabditis elegans. iScience. 2019 Dec 20;22:466-476. doi: 10.1016/j.isci.2019.11.039.
Kaiser RWJ, Ignarski M, Van Nostrand EL, Frese CK, Jain M, Cukoski S, Heinen H, Schaechter M, Seufert L, Bunte K, Frommolt P, Keller P, Helm M, Bohl K, Höhne M, Schermer B, Benzing T, Höpker K, Dieterich C, Yeo GW, Müller RU, Fabretti F. A protein-RNA interaction atlas of the ribosome biogenesis factor AATF. Sci Rep. 2019 Jul 30;9(1):11071. doi: 10.1038/s41598-019-47552-3.
Ignarski M, Islam R, Müller RU. Long Non-Coding RNAs in Kidney Disease. Int J Mol Sci. 2019 Jul 3;20(13):3276. doi: 10.3390/ijms20133276.
Ignarski M, Rill C, Kaiser RWJ, Kaldirim M, Neuhaus R, Esmaillie R, Li X, Klein C, Bohl K, Petersen M, Frese CK, Höhne M, Atanassov I, Rinschen MM, Höpker K, Schermer B, Benzing T, Dieterich C, Fabretti F, Müller RU. The RNA-Protein Interactome of Differentiated Kidney Tubular Epithelial Cells. J Am Soc Nephrol. 2019 Apr;30(4):564-576. doi: 10.1681/ASN.2018090914.
